
Victor Rojas-Zumaran
First Prize

Maria Rocio Lozada Diaz
Second Prize

Harika Derya Tamer
Third Prize
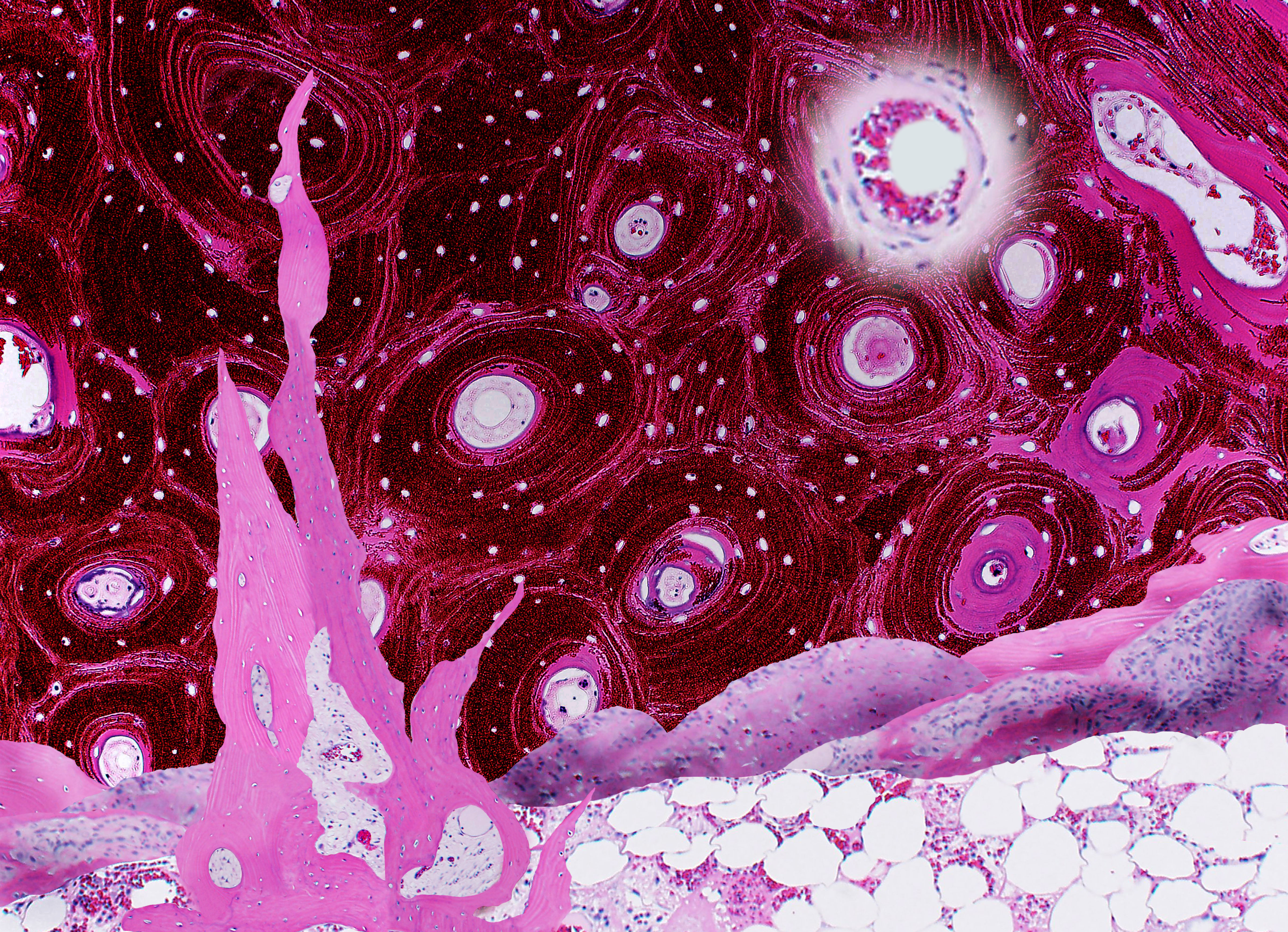
Erica Worswick, USA
First Prize

Esma Kara, Turkey
Second Prize
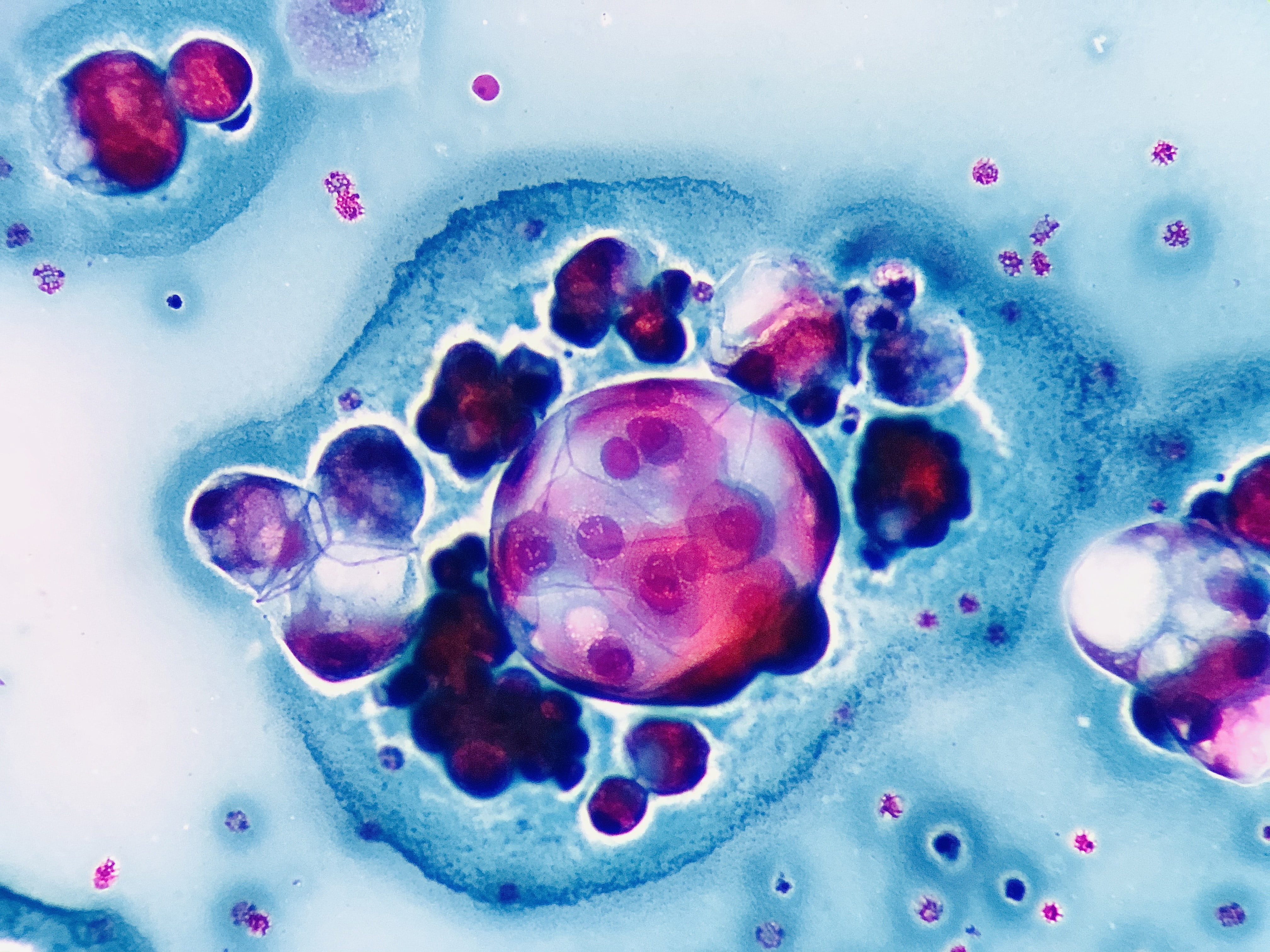
Julio Maciel Mercado, Mexico
Third Prize

Orhan Semerci, Turkey
First Prize

Ishant Anand, India
Second Prize
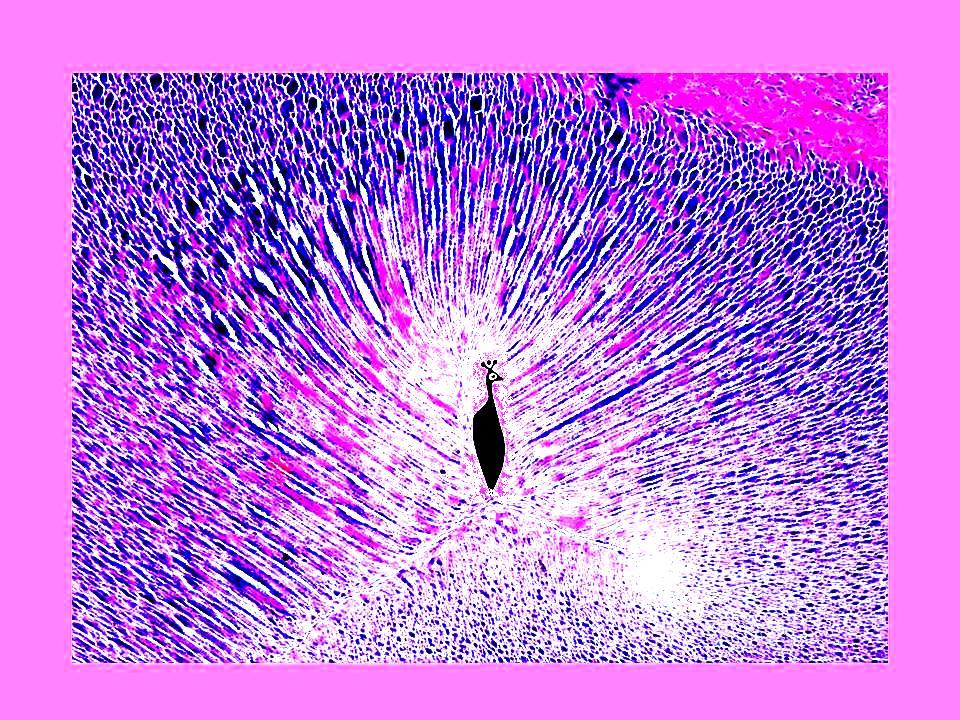

Teodora Tóth, Hungary
Third Prize
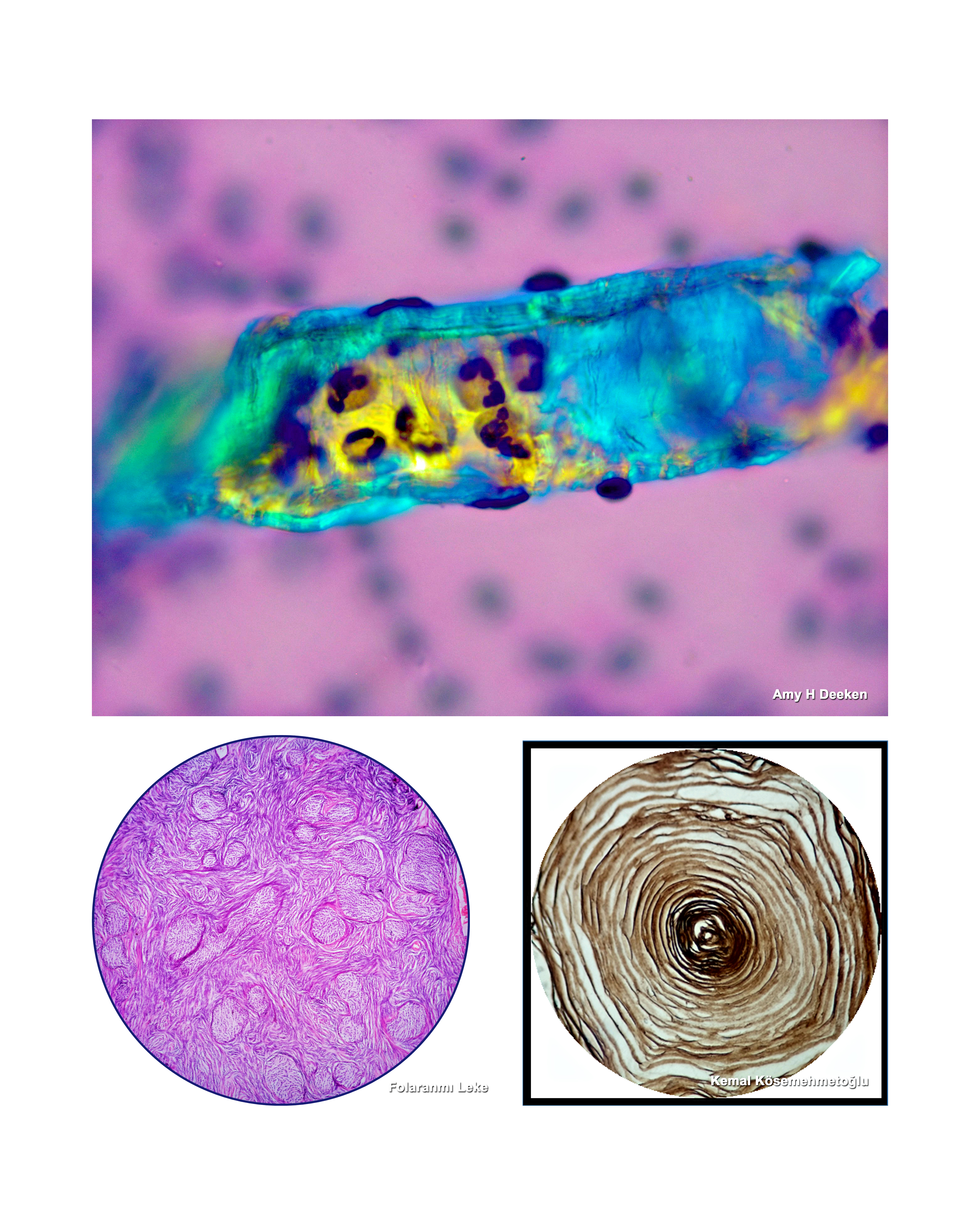
Kayhan Başak
First Prize

Luis Humberto Cruz Contreras
Second Prize
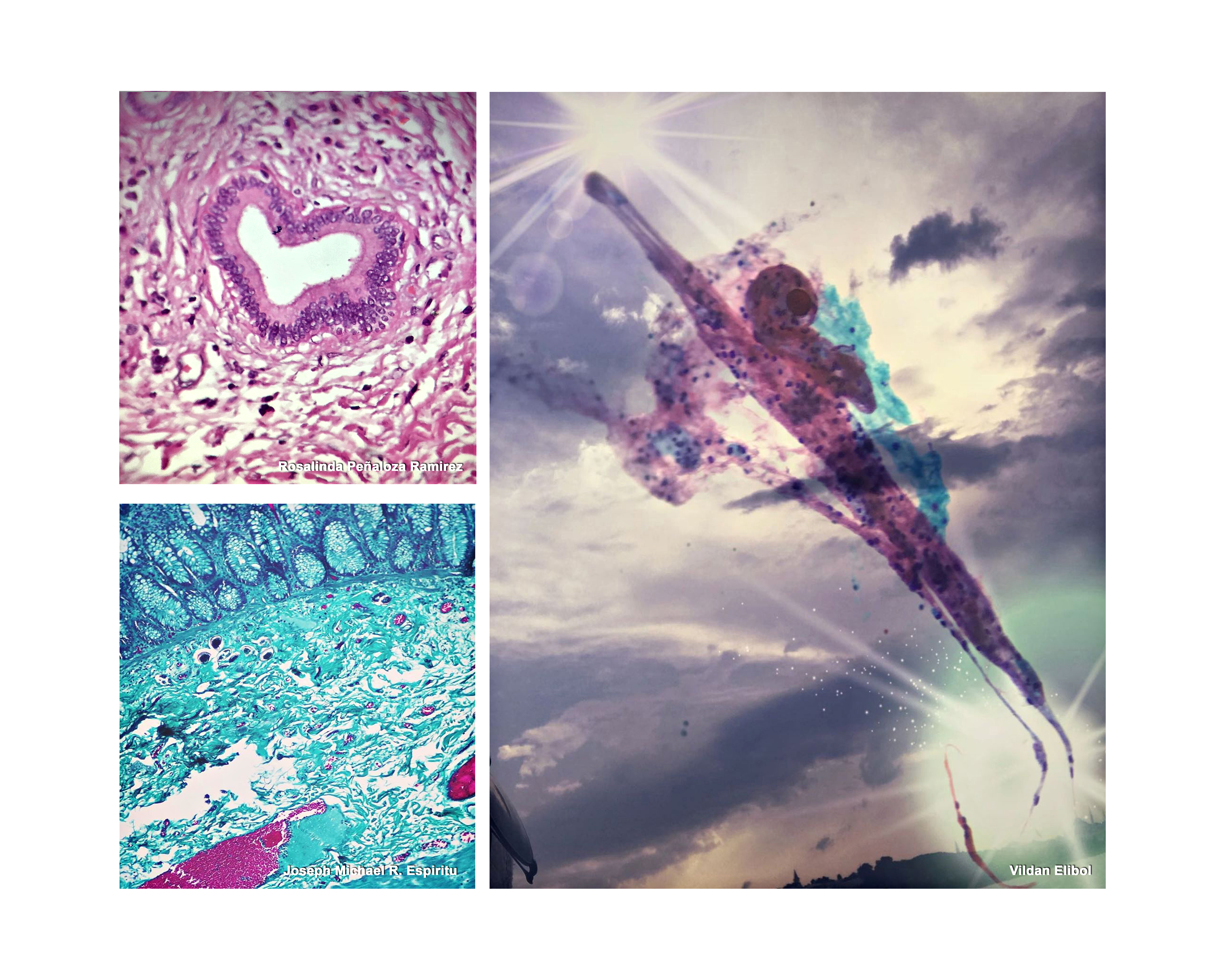
Linda Johnston
Third Prize

Rosalinda Peñaloza Ramirez, Mexico
First Prize
Kayhan Basak, Turkey
Second Prize
Nejib Ben Yahia, Tunisia
Third Prize
